In precision medicine, one of the most persistent challenges is simple yet costly: the same drug can produce dramatically different outcomes in different patients. One patient recovers fully after a standard dose of clopidogrel following cardiac stenting while another individual suffers from a dangerous clot because they carry CYP2C19 which leads to loss of function variants. Adverse drug reactions (ADRs) remain a leading cause of hospitalisations and deaths worldwide. Traditional systems that help in prescribing the drugs are inefficient, expensive, and sometimes dangerous.
Pharmacogenomics (PGx), the study of how genetic variation influences drug response, offers a powerful solution. Yet despite falling sequencing costs and growing availability of genomic data, most hospitals and clinics still struggle to turn genetic insights into routine clinical decisions.
The missing piece is not the science. It's the data engineering.
This blog details how we built a pharmacogenomics clinical decision accelerator on the Databricks Lakehouse platform, a scalable, governed, and standard pipeline that converts raw genotype data into actionable prescribing recommendations at population scale.
The Core Problem
Clinical teams make prescribing decisions every day with limited visibility into a patient's genetic profile. Key challenges include the following:
- Genomic data is complex (VCF files, star alleles, diplotypes, phased variants) and lives in silos.
- CPIC (Clinical Pharmacogenetics Implementation Consortium) and PharmGKB guidelines exist but require manual interpretation, impossible to scale across thousands of patients.
- Prescription workflows in EHR systems are rarely integrated with genomic results.
- Lack of standardised, automated pipelines leads to inconsistent application of evidence-based recommendations.
Solution: The Pharmacogenomics Clinical Decision Accelerator
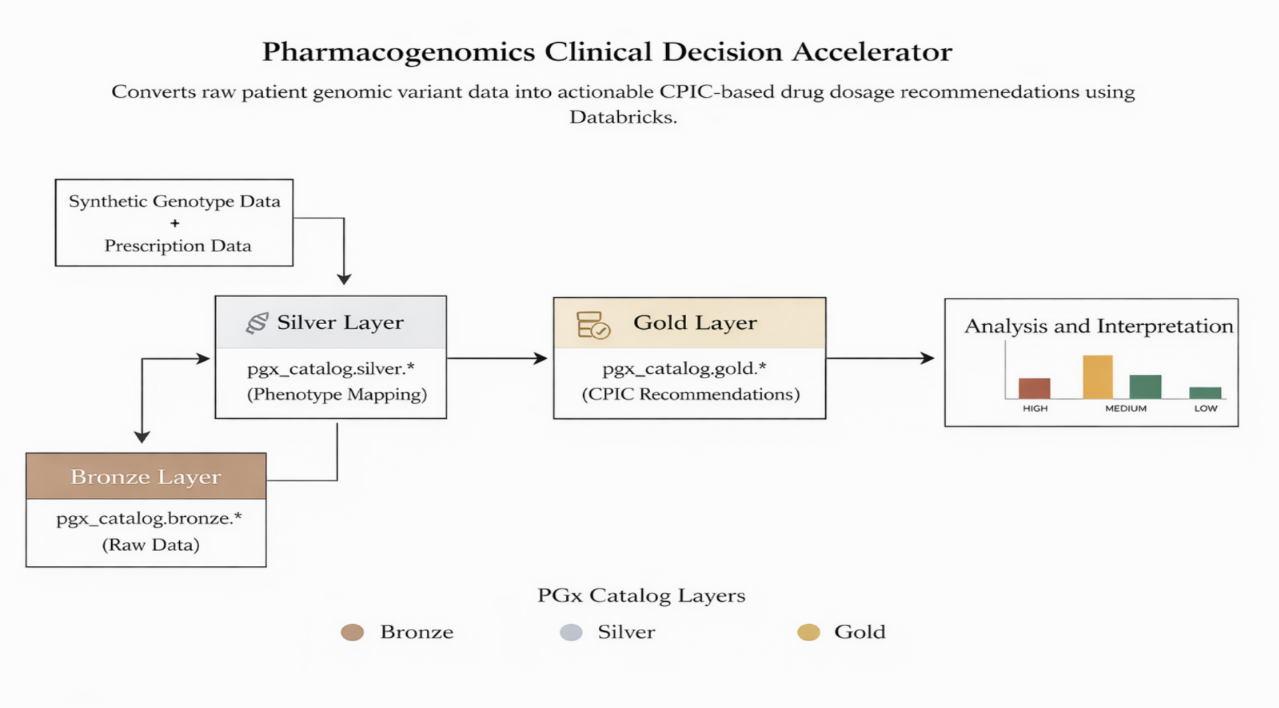
Built on Databricks Lakehouse, the accelerator uses a clean Medallion architecture to ensure reliability, auditability, and scalability while maintaining clinical credibility.
High-Level Architecture:
.png)
.png)
.png)
This layered approach delivers:
- Full data lineage and reproducibility.
- Modular design for easy extension.
- Strong governance through Unity Catalog.
- Scalability ranges from hundreds to millions of patient records.
CPIC-based Integration & Supported Drug-Gene Pairs
We anchored the solution in CPIC guidelines, the internationally recognised clinical standard on the concept covering pharmacogenomics implementation. CPIC provides clear, evidence-based recommendations that translate phenotypes into prescribing actions. Currently supported high-impact pairs include the following:
- Clopidogrel + CYP2C19: Critical for antiplatelet therapy post-stenting.
- Codeine + CYP2D6: Risk of toxicity in ultra-rapid metabolisers or lack of efficacy in poor metabolisers.
- Simvastatin + SLCO1B1: Reduced risk of statin-induced myopathy.
- Warfarin + CYP2C9/VKORC1: Improved dosing accuracy to reduce bleeding or clotting risk.
The given pairs were selected because they have strong Level A or B CPIC evidence and considerable real-world clinical impact.
Key Data Engineering Design Decisions
- PySpark + Delta Lake for distributed processing and ACID compliance.
- Unity Catalog for governance, metadata tagging, and fine-grained access control.
- Explicit schema enforcement and typed UDFs to prevent runtime surprises.
- Avoidance of DBFS limitations by using full catalog tables.
These choices help us ensure the pipeline is production-ready for regulated healthcare environments.
Governance & Compliance
We all know that healthcare data demands rigorous governance. Unity Catalog enables the following:
- Table-level and column-level access controls.
- Sensitivity tagging for PHI/PII.
- Comprehensive audit logging.
- Data lineage tracking.
Delta Lake time travel further supports audit and reproducibility requirements common in clinical systems.
Analytics & Insights
Beyond the patient-level recommendations, the gold layer also provides population-level analytics:
- Phenotype frequency distributions across genes.
- High-risk drug-patient cohort identification.
- Risk heatmaps for drugs and genes.
- Trends in actionable variants across populations.
These insights help health systems identify priority cohorts for PGx testing and measure program impact.
Sample Output: Patient-Level Recommendations
The final deliverable is a clean, queryable table that clinicians or CDS systems can consume:
| Patient_ID | Gene | Phenotype | Drug | Recommendation | Risk | Explanation |
|---|---|---|---|---|---|---|
| P12345 | CYP2C19 | Poor Metabolizer | Clopidogrel | Avoid / Alternative | High | Increased risk of thrombotic events |
| P67890 | CYP2D6 | Ultra-rapid | Codeine | Avoid | High | Risk of morphine toxicity |
This structured output bridges the gap between the genomics labs and point of care for decision making.
Business & Clinical Impact
- Healthcare providers: Reduced adverse drug reactions, fewer readmissions, and more confident prescribing for the people at feasible prices.
- Pharmaceutical companies: The given system provides better patient stratification in clinical trials and post-marketing surveillance.
- Researchers: A scalable platform for population-level PGx studies.
Technical Challenges & Solutions
- Ensuring all the guidelines while handling complex, multi-allelic variant mappings accurately.
- Keeping clinical logic synchronised with evolving CPIC guidelines.
- Ensuring performance at scale without sacrificing precision.
- Maintaining explainability for clinical trust.
Solutions included modular mapping tables, typed transformations, incremental processing patterns, and strong emphasis on auditability.
Future Roadmap
The provided architecture has been designed for growth:
- Add new gene-drug pairs via configuration updates.
- Ingest real VCF files using Auto Loader for streaming pipelines.
- Integrate everything directly with EHR systems for bidirectional data flow.
- Layer in LLM-generated plain-language explanations for clinicians.
- Expand to full clinical decision support (CDS) hooks.
Conclusion
Precision medicine will only succeed if we can operationalise genomic insights at scale. During today's time, using the given technology along with the publicly available data we can combine scalable infrastructure, rigorous governance, and established clinical standards like CPIC, we move healthcare from reactive trial and error towards proactive, personalised care.
Tech Stack
- Platform: Databricks Lakehouse (Unity Catalog, Delta Lake, PySpark)
- Clinical Standards: CPIC Guidelines, PharmGKB
- Processing: Typed UDFs, table-driven rules, Medallion architecture
From raw variants in the bronze layer to clinically actionable recommendations in gold, this Pharmacogenomics Clinical Decision Accelerator shows what's possible when great data engineering meets precision medicine. The future of healthcare isn't just more genomic data—it's making that data usable at the point of care.
Bridging the gap between data and decisions in precision medicine starts with robust, governed, and scalable pipelines.